News from the Institute

ENABLE and SFB 1177 are organizing the 4th Frankfurt Conference on Quality Control in Life Processes (FCQC) which will take place from March 24th – 27th 2025 at Campus Westend of Goethe University Frankfurt.
... (read more)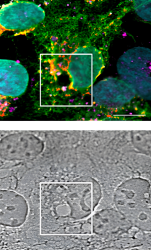
Based on a previous screen in which key kinases of ER-phagy/reticulophagy regulation were identified, the team of IBC2 and BMLS group leader Alexandra Stolz now reveals further insight into the specific roles of two kinases: ATR and CSNK2 (also known as CK2), which both significantly impact ER-phagy flux and act downstream of autophagy master switch MTOR. ER-phagy, or the selective degradation of the endoplasmic reticulum (ER), is vital for cellular homeostasis.
... (read more)
The PROXIDRUGS consortium was successfully evaluated by an independent jury and will be funded by the Federal Ministry of Education and Research (BMBF) as part of Clusters4Future initiative with up to €15 million over the next three years, starting in January 2025. PROXIDRUGS, one of seven initially successful clusters, is thus entering its second implementation phase. A primary aim of the Clusters4Future initiative is to accelerate the transfer of basic research into application.
... (read more)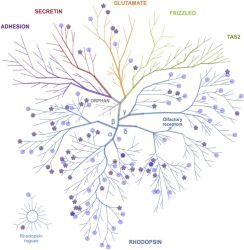
An international team around IBC2 and BMLS group leader Alexandra Stolz just published a study in Journal of Molecular Biology that sheds light on signaling routes that activate individual autophagy pathways. Utilizing a well-characterized chemical library, they systematically screened connections between G-protein-coupled receptors (GPCRs) and autophagy, a crucial cellular process involved in maintaining cell health and responding to stress. Autophagy helps degrade and recycle cellular components, and its dysfunction is linked to various diseases, including cancer, cardiomyopathy and metabolic disorders.
... (read more)
IBC2 Director Ivan Đikić ranked #7 in Germany and #141 in the world on Research.com’s 2024 list of ‚Best Molecular Biology Scientists‘. It is inspiring to see him recognized among this group of pioneering researchers in the field.
... (read more)